Recently, Prof. Donal O’Carroll’s group at the University of Edinburgh (UoE), together with Prof. Wanlu Liu (刘琬璐)’s group at ZJU-UoE Institute (ZJE) and Prof. Elly M. Tanaka’s group at the Research Institute of Molecular Pathology (IMP) in Vienna, published a collaborative study in The EMBO Journal titled “A mammalian-like piRNA pathway in Axolotl reveals the origins of piRNA-directed DNA methylation.” Using the axolotl as a model organism, the study uncovers for the first time the evolutionary origins of the mammalian nuclear piRNA pathway.
Xinyu Xiang (相欣雨), a dual-degree ZJU-UoE PhD at ZJE, is the first author of this study. Her ZJU supervisor is Prof. Wanlu Liu, and her UoE supervisor is Prof. Donal O’Carroll FRS FRSE (corresponding author). Anni Gao (高安妮), an undergraduate student in the bioinformatics dual-degree program at ZJE, is the second author; she conducted research training in Prof. Liu’s lab during her undergraduate studies. This work highlights the strong strengths of the ZJU-UoE Institute in fostering high-level research collaboration and talent development, and exemplifies the impactful synergy and innovation arising from the international partnership between the two universities in the field of biomedical research.

(1) piRNA-mediated transposon silencing is essential for germline integrity
During the development of animal germ cells (such as sperm and oocytes), active transposonsbehave like “genomic time bombs.” As highly abundant mobile DNA elements, transposons can “jump” to new genomic locations, disrupt genes, induce DNA double-strand breaks, and consequently impair germ cell development and lead to infertility. The piRNA (PIWI-interacting RNA) pathwayfunctions as an efficient “demining system,” precisely recognizing and silencing active transposons 【1, 2】. It serves as a critical defense mechanism that safeguards genome integrity in the germline.
(2) The nuclear piRNA pathway is believed to be bespoke to mammals
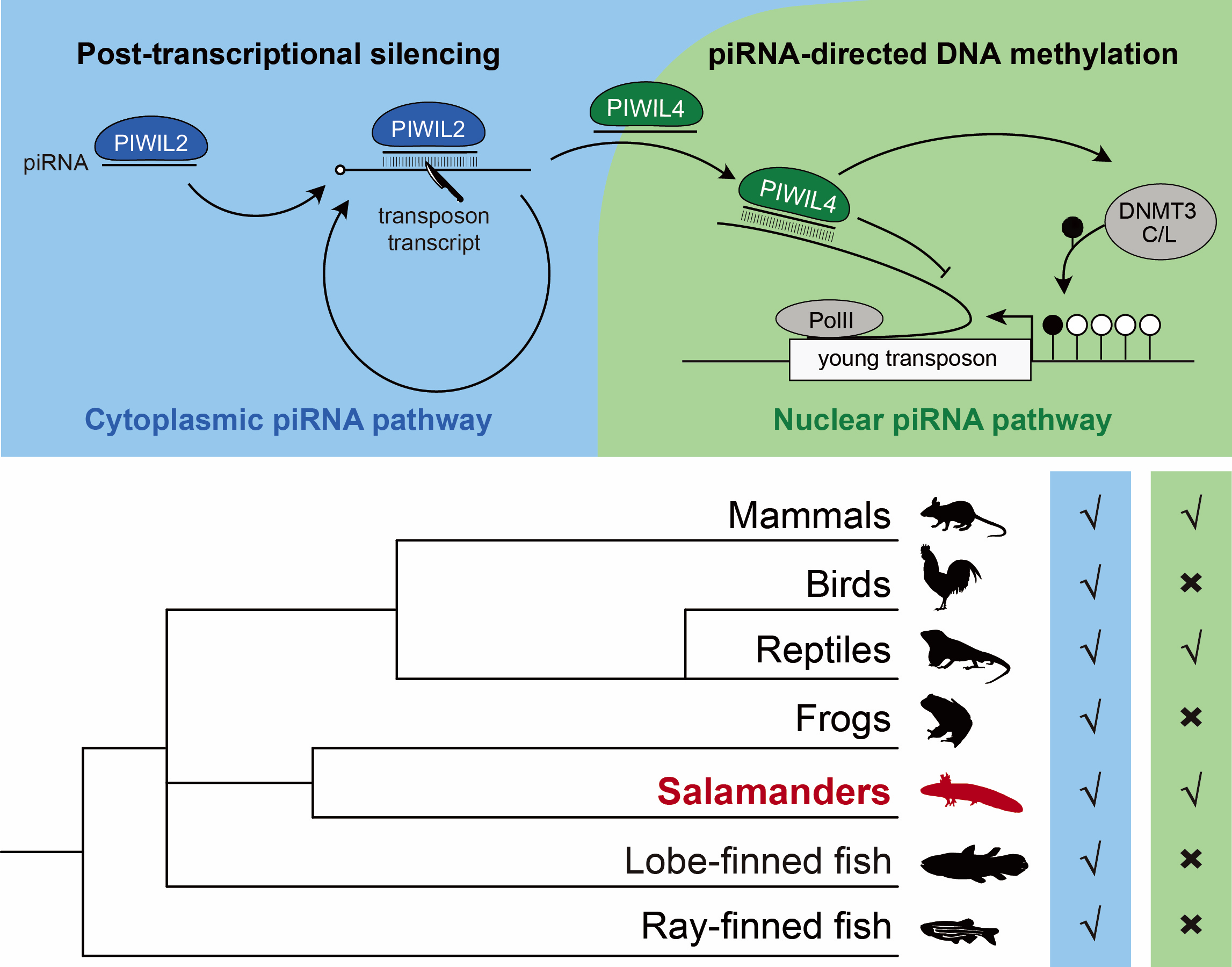
piRNAs are small non-coding RNAs that associate with PIWI proteins to silence transposons in germ cells, effectively defusing these “genomic time bombs.” The piRNA pathway consists of two parts: the cytoplasmic pathwayand the nuclear pathway. The cytoplasmic piRNA pathway is common to most animals 【3】 and functions by cleaving transposon transcripts, providing rapid and timely silencing. In contrast, the nuclear piRNA pathway acts as a “permanent seal,” guiding DNA methylation to stably silence transposons over the long term. For many years, this nuclear piRNA pathway was thought to be a mammal-specific mechanism 【1,2】.
(3) Axolotl is an ideal model for studying transposons and the piRNA pathway
The axolotl is a classic model organism in regenerative biology, well known for its remarkable tissue repair capacity, such as limb regeneration. Beyond its remarkable regenerative abilities, the axolotl also possesses an exceptionally large genome, as one of the largest among known animals (approximately 32 Gb, about ten times the size of the human genome) 【4】. This huge genome is largely result from extensive transposon expansion. Around 70% of the axolotl genome consists of transposons, compared with about 50% in humans. Its high transposon content and huge genome make the axolotl an ideal model for investigating transposon activity and their silencing mechanisms.

(4) Axolotl has a mammalian-like piRNA pathway
To trace the evolutionary origin of the mammalian piRNA pathway, we constructed a phylogenetic tree of key piRNA pathway genes across vertebrates. The analysis revealed that the core nuclear piRNA factors in mammals, such as PIWIL4, SPOCD1, C19ORF84, and TEX15, are found all together for the first time in the axolotl. Whereas these factors are partially missing in both lobe-finned and ray-finned fish. This suggests that the nuclear piRNA pathway may not have emerged in earlier vertebrate lineages. Furthermore, transcriptomic analyses show that these piRNA pathway genes exhibit germline-specific expressionin the axolotl, same as the mammalian expression pattern, thereby indicating their potential functional roles.
(5) Axolotl possesses a functional cytoplasmic piRNA pathway
To better understand transposon activity and silencing in the axolotl, we first predicted and annotated active transposons in axolotl genome, identifying a set of potentially active transposons. We then performed small RNA sequencing to profile piRNAs in axolotl germlines. The results revealed an abundantrepertoire of piRNAs in both male and female germlines (~88 million), with a substantial fraction derived from transposon sequences.
The cytoplasmic piRNA pathway involves piRNA biogenesis and the classic ping-pong cycle, which mediates post-transcriptional silencing by cleaving transposon transcripts. These piRNAs are primarily generated from genomic piRNA clusters, which display clear sex-specific patterns in the axolotl. Moreover, transposon-targeting piRNAs exhibit hallmark features of the ping-pong cycle, providing strong evidences that a functional cytoplasmic piRNA pathway in the axolotl germline.
(6) Evidence for an active nuclear piRNA pathway in axolotl is presented
The nuclear piRNA pathway, or piRNA-directed DNA methylation, has been reported exclusively in mammals. This pathway is mediated by the piRNA-PIWIL4 complex together with associated nuclear factors, which recruit de novo DNA methyltransferases to specifically methylate the promoters of young and active transposons, ensuring their long-term silencing. During epigenetic reprogramming of the germline, the genome undergoes extensive DNA demethylation and is therefore particularly vulnerable to transposon activation, making piRNA-directed methylation silencing especially critical at this stage. The terminal outcome of this process is the high methylation levels of young transposons in sperm.
Whole-genome bisulfite sequencing of axolotl sperm revealed an overall CpG methylation level of approximately 90%, higher than mammals (~75%-85%). Importantly, the entire transposon region, including promoter regions, exhibited tight methylation control. Moreover, young and potentially active transposons also showed significantly higherDNA methylation levels. Together, these findings provide the first evidence outside mammals that DNA methylation mediated transposon silencing, characteristic of the nuclear piRNA pathway, also exists in the axolotl.
Overall, this study identifies a mammalian-like cytoplasmic and nuclear piRNA pathway in the axolotl. Small RNA sequencing and whole-genome methylation profiling together demonstrate that the axolotl possesses a functional cytoplasmic piRNA pathway for transposon transcript cleavage, as well as a nuclear piRNA pathway for long-term transposon silencing through DNA methylation. This work provides the first evidence that the mammalian piRNA pathway, particularly the piRNA-directed DNA methylation mechanism, can be traced back to the common ancestor of the axolotl and mammals, which are the early tetrapods. These findings not only fill a critical gap in the evolution of the piRNA pathway but also offer new insights into germline development, epigenetic reprogramming, and genome defense mechanisms.
Original Publication:
https://www.embopress.org/doi/full/10.1038/s44318-025-00631-w
【1】 Wang X, Ramat A, Simonelig M, and Liu M-F Emerging roles and functional mechanisms of PIWI-interacting RNAs Nat Rev Mol Cell Biol 2023 24 123-141
【2】 Ozata DM, Gainetdinov I, Zoch A, O’Carroll D, and Zamore PD PIWI-interacting RNAs: small RNAs with big functions Nat Rev Genet 2019 20 89-108
【3】 Grimson A, Srivastava M, Fahey B, Woodcroft BJ, Chiang HR, King N, Degnan BM, Rokhsar DS, and Bartel DP Early origins and evolution of microRNAs and Piwi-interacting RNAs in animals Nature 2008 455 1193-1197
【4】 Nowoshilow S, Schloissnig S, Fei J-F, Dahl A, Pang AWC, Pippel M, Winkler S, Hastie AR, Young G, Roscito JG, et al. The axolotl genome and the evolution of key tissue formation regulators Nature 2018 554 50-55







